Help - How to use GASS-Metal
Main page
About GASS-Metal
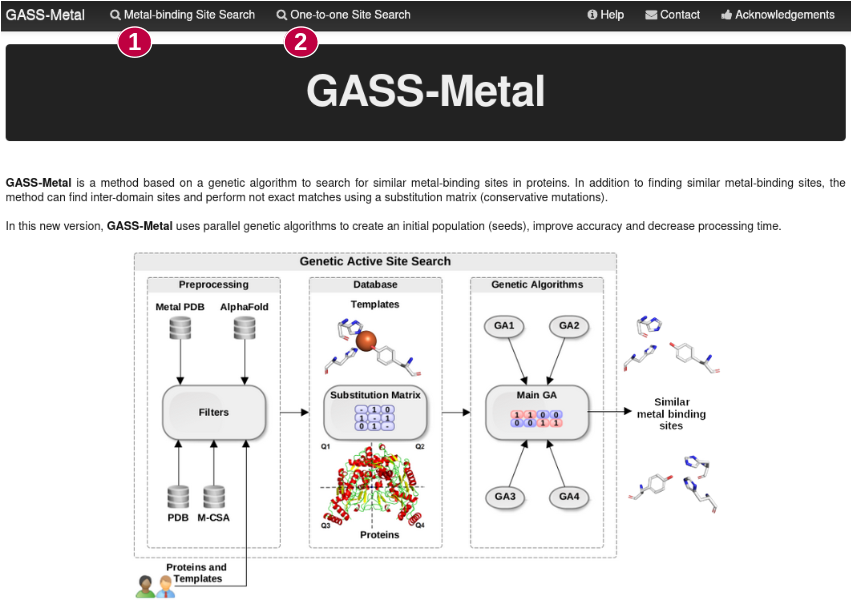
GASS-Metal is a user-friendly web server based on parallel genetic algorithms to predict
metal-binding sites in protein structures, based on similarity of knows sites. It has two available resources:
-
Metal-binding Site Search (1): Given a protein of interest, try to match a metal site based on M-CSA and MetalPDB.
-
One-to-one Site Search (2): The user can give the protein target and your template.
Metal-binding Site Search
How to run Metal-binding Site Search
To perform a site search based on M-CSA and MetalPDB:
-
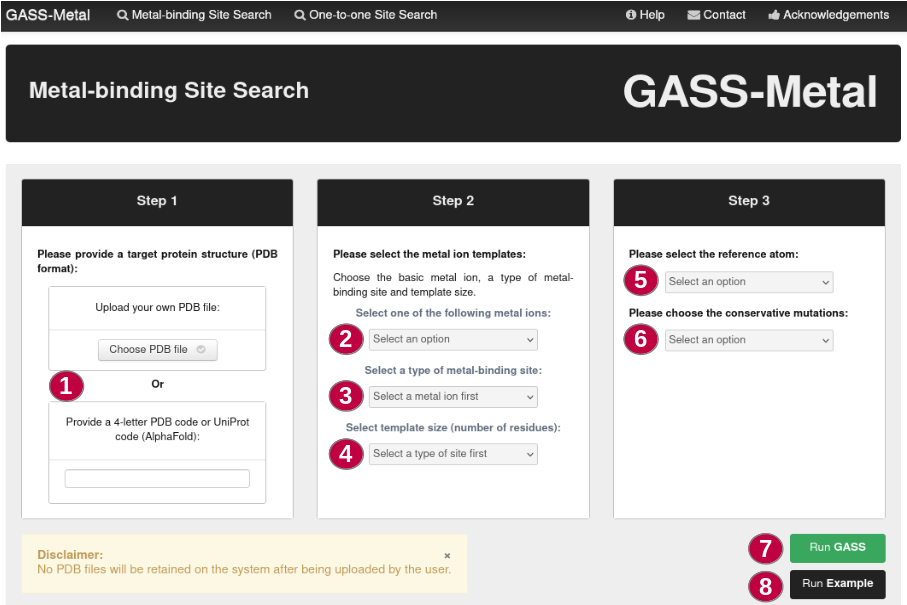
Step 1: The user must provide a 4-letter PDB code or UniProt code (AlphaFold), or provide a PDB file (1), which will serve as a target for searching metal-binding sites. It is in this protein that GASS-Metal will search for similar sites using its templates.
-
Step 2: Where the templates on which the search for GASS-Metal will be based are filtered. Users must first choose the metallic ion of the site (2), the type of this site (if it is simple, containing only one ion, or compound, having two or more metal ions) (3) and the number of residues that make up the templates (4). The All option enables GASS-Metal to perfom an exploratory search considering all templates for that metal ion.
-
Step 3: The user can choose between the alpha carbon or the last heavy atom (LHA) as the reference atom in the templates (5). The user also can choose one of three options about conservative mutations (6): use mutations provided by GASS-Metal, use user-defined mutations or use no mutations at all.
The button Run Example (8) will perform a metal-binding site search based on a pre-defined example.
Processing Page
Processing Page
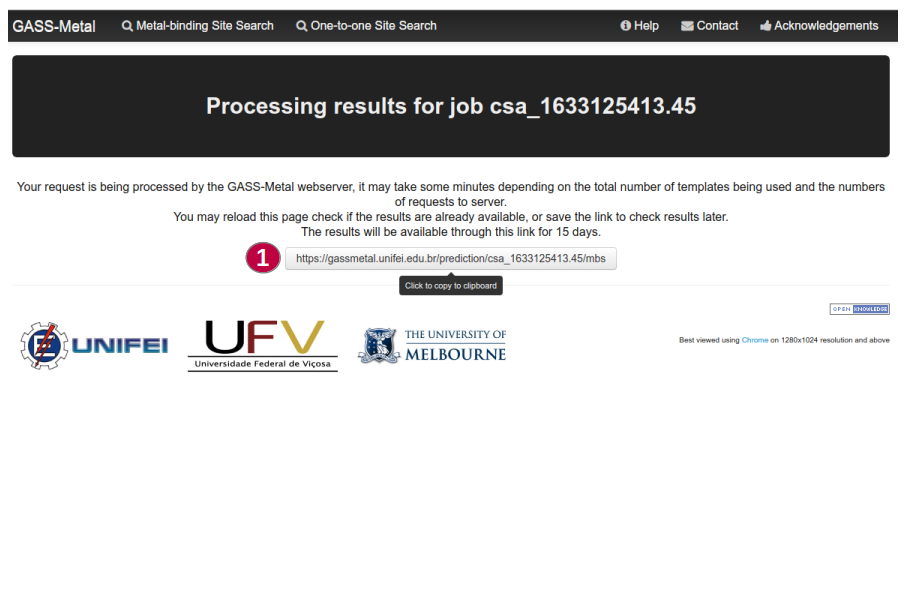
A request made to the GASS-Metal web server may take a few minutes depending on the total
number of templates being used and server load.
Users may reload the results page to check if the results are already available or save the
link to check results later (1).
The results will be available through the link for 15 days.
Results page
Results
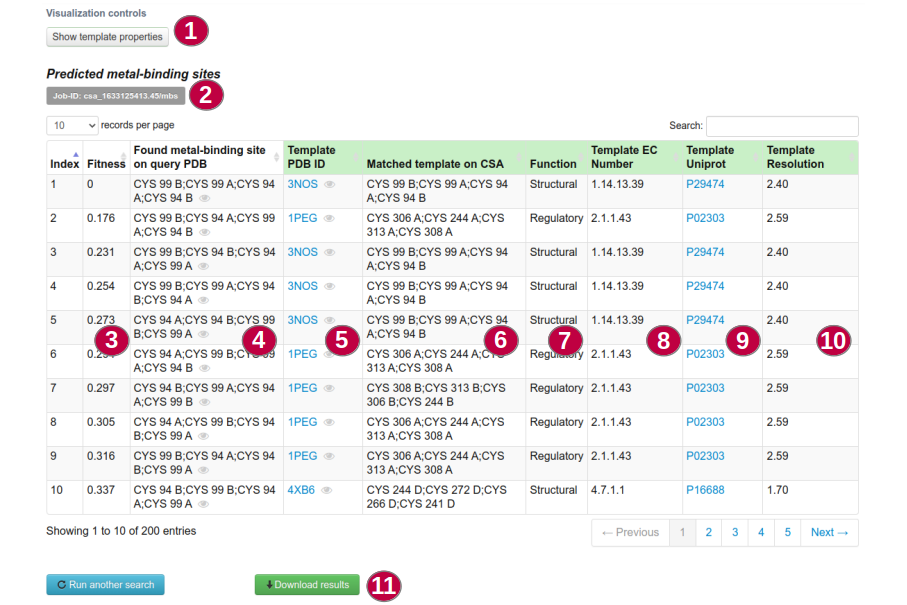
In Metal-binding Site Search, your results will be displayed in a table format with the following information:
-
Shows or hides options and properties of templates (1).
-
Job ID (2) - can be used to retrieve the results later adding it to the webserver URL following the format:
http://gassmetal.unifei.edu.br/prediction/csa_1633125413.45/mbs
-
Fitness score of matched residues (3).
-
Residues found by GASS-Metal on input structure matching a template(4).
-
PDB ID of a matched template (5).
-
Residues of matched template (6).
-
Function of the metal site (7), EC Number (8), Uniprot accession code (9) and Resolution (10) of a matched template.
-
The results may also be downloaded as a CSV file (11).
-
The icon in the shape of an eye (4 and 5) opens the LiteMol Viewer.
LiteMol Viewer
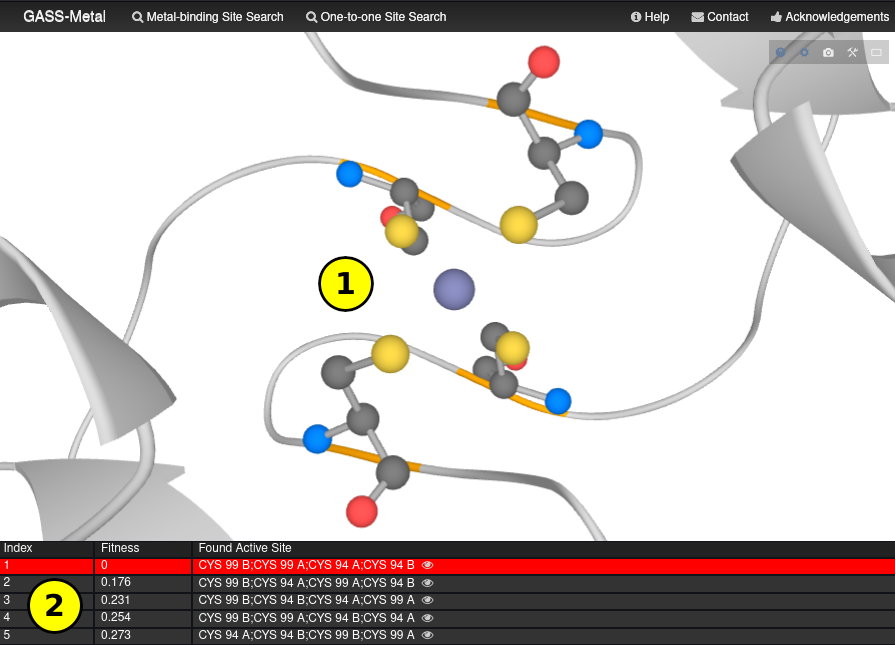
LiteMol Viewer
The amino acid residues found by GASS-Metal can be visualized through the icon in the shape of an eye in the result pages.
This icon starts LiteMol and displays the amino acid residues found in stick representation and cartoon representation highlighted in orange (1).
The GASS-Metal results list is also available (2).
One-to-one Site Search
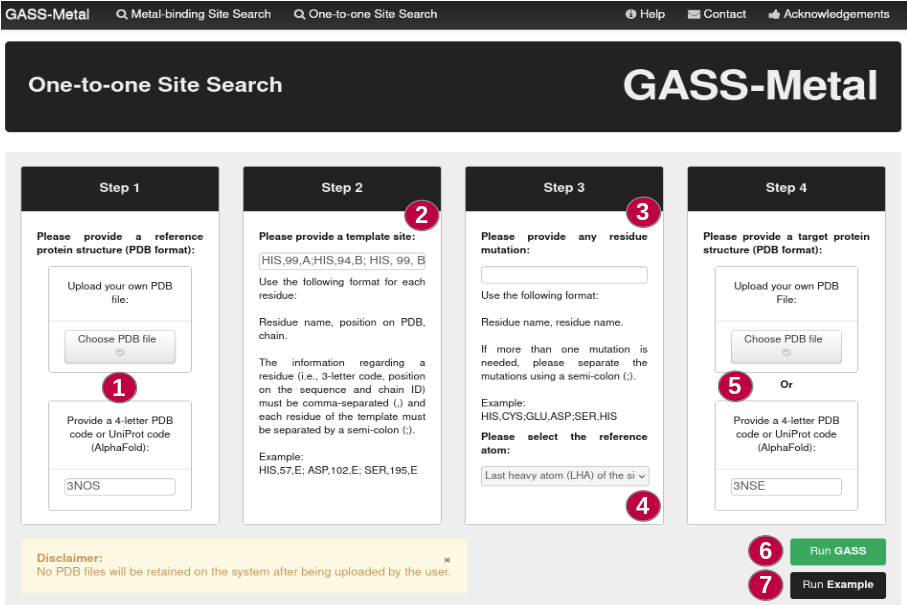
How to run One-to-one Site Search
To perform a site search based on your own template:
-
Step 1: Users must provide a 4-letter PDB code or UniProt code (AlphaFold), or provide a PDB file (1), which will serve as a reference for searching metal-binding sites by GASS-Metal.
-
Step 2: Once the protein has been defined, it must be indicated what the reference template (which must be contained in the reference protein) on which the search will be based (2).
-
Step 3: In this step, users can indicate whether to use conservative mutations or not (3). When choosing conservative mutations, inform which residues can be replaced in the target protein. Users can choose between alpha carbon or the last heavy atom (LHA) as the reference atom in the templates (4).
-
Step 4: Users must provide a 4-letter PDB code or UniProt code (AlphaFold), or provide a PDB file (5), which will serve as a target for searching metal-binding sites. It is in this protein that GASS-Metal will search for sites similar to the template informed in step 2.
The button Run Example (7) will perform a metal-binding site search based on a pre-defined example.